Genes Expression And Its Regulation
Category : 12th Class
Gene expression in prokaryotes
Gene expression refers to the molecular mechanism by which a gene expresses a phenotype by synthesizing a protein or an enzyme. Which determines the character. The gene contains the blue print or the information for the protein or an enzyme.
The category includes mechanism involved in the rapid turn-on and turn-off gene expression in response to environmental changes. Regulatory mechanism of this type are very important in microorganisms, because of the frequent exposure of these organisms to sudden changes in environment.
Gene concept can be studied by operon model. Operon are segment of genetic material which function as regulated unit that can be switched on and switched off, which was given by French scientists. Jacob and Monod (1961) working at Pasteur institute. They were studying lactose utilization in mutants of E.coli. An operon consists of one to several structural genes (three in lac operon and five in tryptophan operon of Escherichia coli, nine in histidine operon of Salmonella typhimurium), an operator gene a promoter gene a regulator gene, a repressor and inducer or corepressor. Operons are of two types, inducible and repressible.
(1) Inducible operon system /lac operon system : An inducible operon system is that regulated genetic material which remains switched off normally but becomes operational in the presence of an inducer. It occurs in catabolic pathways. The components are:
Structural genes : They are genes, which produce mRNAs for forming polypeptides/proteins/enzymes. Lac operon of Escherichia coli has three structural genes-Z (produces enzyme \[\beta -\]galactosidase for splitting lactose/galactoside in to glucose and galactose) Y (produces enzyme galactoside permease required in entry of lactose/galactoside) and A (produces enzyme galactoside acetylase/transacetylase without any function in E.coli). The three structural genes of lac operon produce a single polycistronic mRNA. The three enzymes are, however, produced in different concentration.
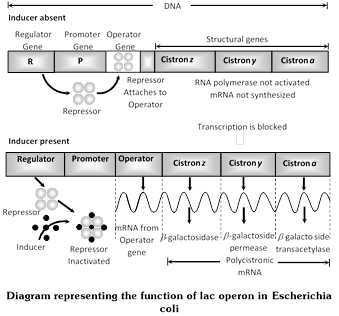
Operator gene (O) : It gives passage to RNA polymerase when the structural genes are to express themselves. Normally, it is covered by a repressor. Operator gene of lac operon is small, made of 27 base pairs.
Promoter gene (P) : It is recognition centre / initiation point for RNA polymerase of the operon.
Regulator gene (i Gene) : It produces a repressor that binds to operator gene for keeping it nonfunctional (preventing RNA polymerase to pass from promoter to structural genes).
Repressor : It is a small protein formed by regulator gene. Which binds to operator gene and blocks passage of RNA polymerase towards structural enzymes. Repressor has two allosteric sites, one for attaching to operator gene and second for binding to inducer. Repressor of lac operon has a molecular weight of 160,000 and 4 subunit of 40,000 each.
Inducer : It is a chemical which attaches to repressor, changes the shape of operator binding site so that repressor no more remain attached to operator.
Differences between induction and repression
|
Induction |
Repression |
|
It turns the operon on. |
It turns the operon off. |
|
It starts transcription and translation. |
It stops transcription and translation. |
|
It is caused by a new metabolite which needs enzymes to get metabolised. |
It is caused by an excess of existing metabolite |
|
It operates in a catabolic pathway. |
It operates in an anabolic pathway. |
|
Repressor is prevented by the inducer from joining the operator gene. |
Aporepressor is enabled by a corepressor to join the operator gene |
Lactose/galactoside is inducer of lac operon. As soon as the operator gene becomes free, RNA polymerase is recognised by promoter gene. cAMP is required, RNA polymerase passes over the operator gene and then reaches the area of structural genes. Here it catalyses transcription of mRNAs.
(2) Repressible operon system/tryptophan operon system : A repressible operon system is that regulated genetic material, which normally remains active/operational and enzymes formed by its structural genes present in the cell till the operon is switched off when concentration of an end product crosses a threshold value. Repressible operon system usually occurs in anabolic pathways, e.g., tryptophan operon, argnine operon. Each has the following parts.
Structural genes : They are genes, which take part in synthesis of polypeptides/proteins/enzymes through the formation of specific mRNAs. Tryptophan operon has five structural genes – E, D, C, B and A.
Operator gene : It provides passage to RNA polymerase moving from promoter to structural genes. Operator gene of repressible operon is normally kept switched on as aporepressor formed by regulator gene is unable to block the gene.
Promoter gene : It is initiation/recognition point for RNA polymerase.
Regulator gene : The gene produces an aporepressor.
Aporepressor : It is a proteinaceous substance formed through the activity of regulator gene. It is able to block operator gene only when a corepressor is also available.
Corepressor : The nonproteinaceous component of repressor, which can be end product (feed back inhibition/repression) of the reaction mediated through enzymes synthesized by structural genes. Corepressor of tryptophan operon is tryptophan. It combines with aporepressor, form repressor which then blocks the operator gene to switch off the operon.
Gene expression in eukaryotes
In regulation of gene expression in eukaryotes the chromosomal proteins play important role. The chromosomal proteins are of two types. They are histones and non-histones. The regulation of gene expression involves an interaction between histones and non-histones. Histones inhibit protein synthesis and non-histones induce RNA synthesis. There are four main steps in the expression of genes. Hence regulation is brought about by the regulation and modification of one or more of these steps. They are :
Regulation of replication : Differential gene expression is achieved by gene amplification.
Regulation of transcription : The regulation of the expression of gene is mainly done at transcription. Hybridization experiments clearly show that production of specialised protein is due to differential gene transcription.
Regulation of the processing level : Some of the RNA synthesized in the nucleus are destroyed without leaving the nucleus. 80% of the nuclear RNA has no equivalent in the cytoplasm and only 20% if the nuclear RNA is identical in the cytoplasm. All the genes in a cell are transcribed into mRNA at all times, but the mRNA produced by some genes is destroyed rapidly. But the mRNA modeled on other genes are stabilized and only these mRNAs are passed into the cytoplasm.
Regulation of translation : The control of mRNA-translation is a fundamental phenomenon. In sea-urchin eggs fertilisation is followed by a tremendous increase in protein synthesis; but in the unfertilised egg, there is no protein synthesis. Still the unfertilised egg has complete machinery (i.e., amino acids, ribosomes, mRNA) for protein synthesis. There are two model for regulation in eukaryotes.
(a) Frenster's model : According to 1965, The histones act as repressor's during protein synthesis.
(b) Britten Davidson model : This is also called gene battery model or operon-operator model. It was proposed by Britten and Davidson in 1969. They have been proposed four type of genes namely integrator sensor, producer and receptor.
You need to login to perform this action.
You will be redirected in
3 sec
